Watch bacteria transmit antibiotic resistance through sex in real time
New research shows microbes can become drug-resistant even while being treated with antibiotics.
Improper use of antibiotics makes it easier for drug-resistant strains of bacteria to arise. But as even drug-sensitive bacteria find loopholes, the problem might be even more dire than once thought. Image Credit: Stas_V, iStock
It started, as many things do, with sex.
Sort of.
Like many microbiologists before her, Sophie Nolivos had built a career studying bacterial “sex”—a process called conjugation, in which a donor cell transfers genetic information to a recipient through physical contact. Though conjugation doesn’t yield offspring, it’s an act that still blends the DNA of two individuals in a single cell, often dramatically reshaping the recipient’s genome.
It’s long been known that bacterial conjugation can hasten the spread of antibiotic resistance. But many of the nitty gritty details of these microscopic transactions, including how quickly newly transferred traits are expressed, remained murky. So four years ago, Nolivos, now at the University of Pau and Pays de l’Adour, and Christian Lesterlin, a microbiologist at the University of Lyon, set out to do what no one had done before: observe and track the sexual transmission of antibiotic resistance in single E. coli cells.
Their research, published today in the journal Science, shows the recipients of bacterial conjugation can acquire immunity to antibiotics in a matter of minutes. It’s a process so speedy that some bacteria can even convert from drug-sensitive to drug-resistant while the cells are being bombarded by the very antibiotics they’re vulnerable to.
This on-the-spot transformation is made possible by a common but mighty microbial superpower: the ability of bacteria to jettison drugs back into the environment via a series of pumps that stud their exteriors. In the wake of conjugation, this stalling tactic is akin to bailing water from a sinking ship, buying microbes the time they need to repair the figurative hole in the hull.
“This paper explores this idea beautifully,” says Monika Avello, a bacteriologist and Digital Learning Lab Fellow at MIT who was not involved in the study. “We know that antibiotics are meant to kill bacteria, and resistance is a big problem...but what this [study] shows is that resistance is so much more complicated than we ever thought.”
Though the findings have yet to be tested outside a laboratory, they suggest a sobering reality in which even presumably antibiotic-sensitive bacteria aren’t guaranteed to fall in the face of properly administered drugs. If the results hold true in a host, they might someday prompt a reevaluation of the ways bacterial infections are addressed in the clinic.
“The dream is to develop [antibiotics] that aren’t easily subverted by the evolution of resistance,” says Manoshi Datta, a microbiologist studying antibiotic resistance at the Technion – Israel Institute of Technology who was not involved in the study. “But bacteria are incredibly resourceful…[this] makes it clear we have a challenging path ahead.”
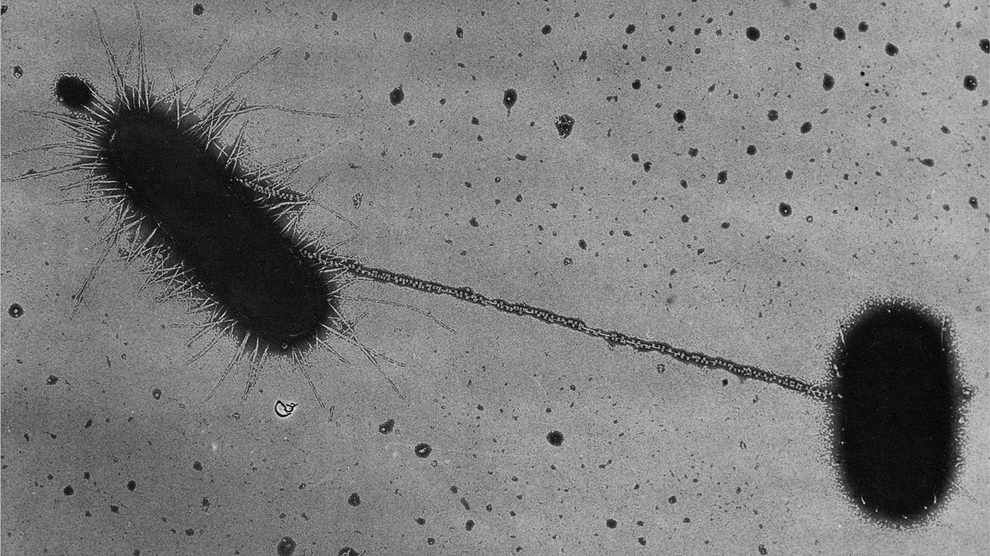
Bacterial conjugation between two E. coli cells. A donor (left) extends a sex pilus to touch the recipient (right) and threads DNA through the tube, passing on traits that can include antibiotic resistance. Image Credit: Charles C. Brinton Jr., NIH
Under ideal conditions, conjugation can be a pretty efficient process, with upwards of 70 percent of recipient bacteria acquiring new genetic material from their donors within minutes of initiating contact. But this high-speed transfer is just the first step on the road to resistance. Even after a conjugation recipient acquires its donor’s DNA, it still needs to read the genes coded within and manufacture proteins before tolerance can manifest.
This delay, which can take hours, has long led researchers to assume that antibiotic-sensitive recipients would need to wrap up any resistance-conferring rendezvous long before being hit with drugs. So when Nolivos mingled drug-resistant donors and drug-sensitive recipients in the presence of tetracycline, she was certain none of the recipients would survive—save for those exceedingly rare individuals that, by chance, already carried a mutation that made them immune to the antibiotic.
But when Nolivos peered into the microscope, she was shocked to see the tetracycline-sensitive E. coli were still alive and kicking. Despite being flooded with antibiotics, the supposedly defenseless recipients were still somehow acquiring genes from donors—and churning out the proteins necessary to resist the drug’s effects.
This was especially baffling, Nolivos says, given that tetracycline specifically targets the cellular machinery responsible for synthesizing proteins. It was like watching a person freshly paralyzed by poison casually whip up and self-administer a home remedy. How was it possible to become immune to a drug that blocked the very process the cells needed to withstand its effects?
The answer turned out to be the product of an educated, and somewhat lucky, guess, Lesterlin says. Because the cells were still conducting business as usual, something in the cell had to be clearing the path to protein production—likely by disposing of the antibiotics themselves.
So-called efflux pumps—which, like human kidneys, play roles in flushing noxious substances out of cells—fit the bill. These detoxifying proteins are widespread among bacteria, and are capable of dumping a variety of antibiotics, including tetracycline, back into the environment. Their role in acquiring antibiotic resistance, however, was something “entirely unknown,” Lesterlin says.
When the researchers deleted one such pump from E. coli, the survival of drug-sensitive recipients plummeted. Conversely, bacteria forced to manufacture pumps in excess were better equipped to acquire tetracycline resistance. The efflux pumps had, in effect, given the microbes the exact loophole they needed to resolve the protein synthesis paradox.
The pumps weren’t enough to confer drug resistance on their own, however, and they didn’t guarantee success. For a lucky subset of bacteria that had acquired the genes necessary for tetracycline resistance, however, these pumps chucked out just enough of the antibiotic to give the cell’s protein assembly line a head start.
And once the process got going, it didn’t take much to cross the threshold to tolerance. “In about 100 minutes, [bacteria] can transfer DNA and do all they need to do to acquire resistance,” Lesterlin says. “Then they do as well as cells that started out resistant.”
Because these experiments were conducted in a laboratory and not in a more “natural” system, there’s no guarantee this process is happening during infections, Datta points out. But, she says, given how dense and diverse microbial communities are, and how widespread these efflux pumps are among bacteria, this represents a “very plausible explanation for how bacteria can acquire [many kinds of] antibiotic resistance.”
If that’s the case, efflux pumps could even be aiding and abetting more than conjugation. Bacteria acquire and develop drug tolerance in a variety of ways, sometimes pilfering genes from viruses, slurping up DNA from their surroundings, or simply accruing spontaneous mutations all on their own. These theories haven’t been tested directly, Lesterlin says, but they’re exciting next steps.
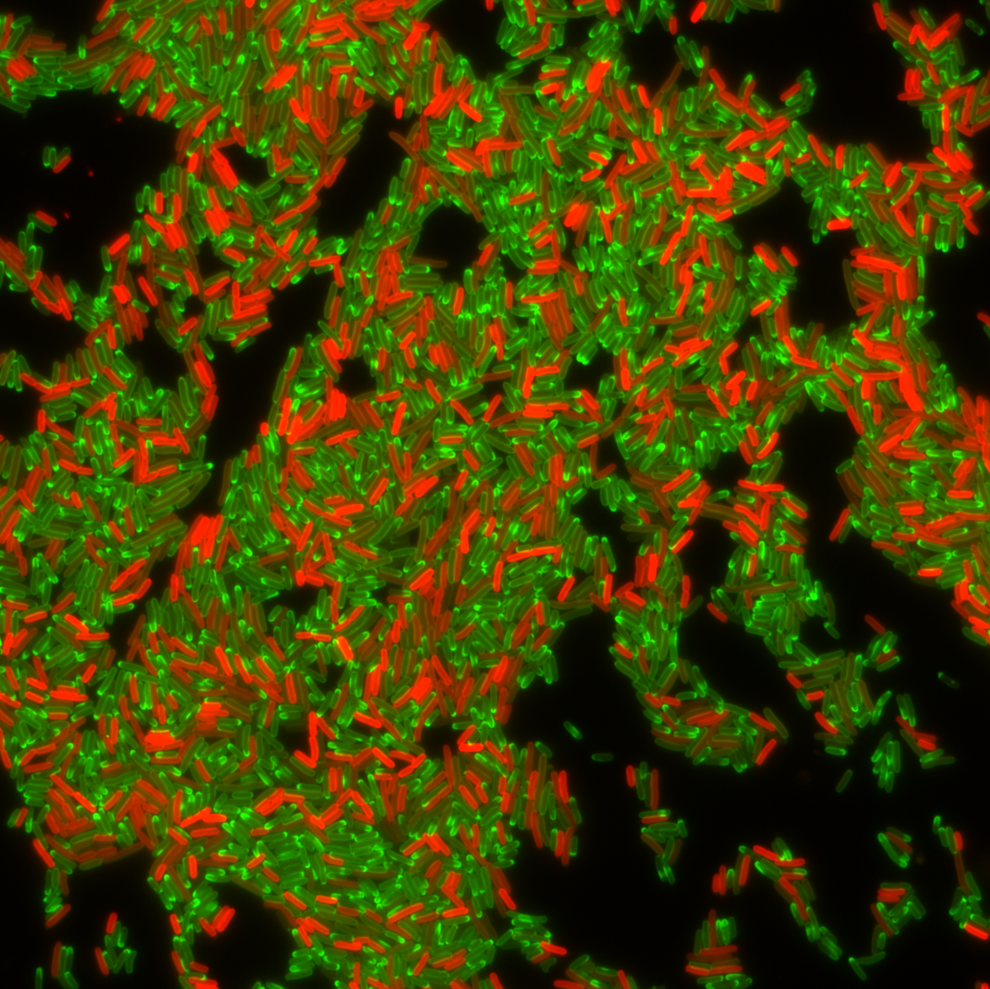
Population of drug resistant bacterial cells producing a protein (red) that confers resistance to the antibiotic tetracycline during treatment with tetracycline (green). Image Credit: Courtesy of Christian Lesterlin, University of Lyon
The study’s findings could also prompt renewed interest in designing drugs to inhibit efflux pumps, he says. Though no such treatments are currently available in the clinic, several pump inhibitors are in development for their potential to increase the sensitivity of bacteria to certain antibiotics, which would accumulate more quickly inside of cells in the absence of pumps. In the future, adding a pump blocker to a traditional antibiotic regimen may have the added perk of stymieing the spread of drug resistance, upping the chances of successful treatment.
All this goes to show “just how many weapons bacteria have to facilitate their evolution and adaptation,” Lesterlin says. Despite their petite size, microbes are as flexible and resilient as can be—and that’s not always a bad thing. Bacteria may give us grief, but we also wouldn’t be here without them.
“[These pumps] are important players in bacterial survival and fitness...they offer flexibility and resilience,” Lesterlin says. “That’s most obviously been beneficial for the bacteria, but sometimes that’s good for us as well.”



